CH391L/S14/SmallRNAs
Contents |
Bacterial small RNAs: as a potential powerful tool for metabolic engineering
Introduction
Bacterial small RNAs (sRNAs) are gene regulatory entities, analogous to their counterparts in eukaryotes micro RNAs, that range from 21 to 400 nucleotides in size. These RNAs are in charge of controlling expression of stress-response genes and thus are essential for an organism's survival under different extreme environmental conditions (e.g. nutrient availability, osmolarity, pH and temperature)[1]. The presence of these regulatory molecules appears to be ubiquitous as they have been discovered in a wide range of bacterial species [2][3]. Their high modularity and orthogonality have raised interest among synthetic biologists towards the construction of sRNA-like devices. In addition, sRNA capacity to simultaneously target single or multiple genes with high specificity has enabled the vision of sRNAs as a powerful tool for metabolic engineering applications.
Bacterial small RNAs

sRNAs can be classified as cis-encoded and trans-encoded. The former refers to those that are transcribed from the complementary strand of the genes that they target. This class represents the minority of the sRNAs that have been identified up to now. Additionally, cis-encoded sRNAs usually exert a tight control over a single target messenger RNA (mRNA). In contrast, trans-encoded sRNAs are transcribed from loci in the genome that are distant from where their mRNA targets are encoded. This class accounts for the great majority of sRNAs discovered to date. An astonishing feature is that these molecules can bind their mRNA partners by a minimal base-pairing requirement (8-9 nucleotides)[1]. Lastly but more importantly, this class of sRNAs can interact with multiple mRNAs[5]. This property, in turn, enables the potential application of combinatorial gene knockdown in metabolic engineering.
Trans-encoded sRNAs can target proteins in addition to mRNAs; an example of that are sRNAs such as CsrB/C and 6S RNA. When controlling mRNA expression this class of sRNAs uses a diversity of mechanisms. They can (1) base-pair to their target mRNAs to enhance or attenuate transcription (Figure 1A), (2) directly block (Figure 1B i), or indirectly enhance or inhibit translation (Figure 1B ii), (3) sequester proteins (not shown), or (4) directly lead to mRNA and protein degradation (Figure 1B iii). This article will exclusively focus on those sRNAs that are trans-encoded and only target mRNAs. Hereafter, they will be referred simply as sRNAs. This class of sRNAs, as aforementioned, accounts for the majority of discovered sRNAs and can target multiple genes. Consequently, these sRNAs have attracted much interest among the Synthetic Biology community as it will be shown in the remainder of this article.
A particular feature that this class of sRNAs exhibits is the interaction with a major chaperone protein called Hfq. These interactions have been mainly observed in gram-negative bacteria. Hfq action leads to the stability of sRNAs, assists their binding to target mRNAs and stabilizes interactions sRNA-mRNA[1]. Recent reports propose that Hfq can also exert negative regulation by delivering the sRNA-mRNA complex to the degradosome [3]. By engineering Hfq interaction, gene expression control could potentially be greatly improved since the gene repression dynamic range is enhanced. In addition, the introduction of Hfq domains into an already constructed sRNA-like device could bring about a very valuable increase in its gene silencing capabilities[6].
sRNAs in Synthetic Biology
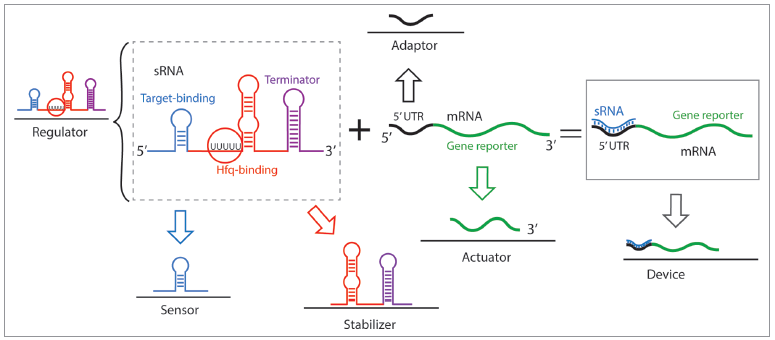
sRNAs are highly composable, (composability is the ability of a system to berak down in units due to the system modularity and recombine in different configurations to satisfy specific human requirements), tunable and their orthogonality can be designed a priori. In general, a variety of strategies have been used to synthesize sRNAs that include rational design, model-driven computational design, in vivo and in vitro molecular evolution and selection and, harvesting of natural parts [4]. Efforts have focused on preserving the sRNA scaffold, which includes an Hfq domain and a transcriptional terminator, and engineering the binding domain (see Figure 2 for a schematics of sRNA breakdown).
Designing a synthetic sRNA

Three factors likely influence sRNAs ability to regulate gene expression: kinetics of binding, extension and energy of binding as well as the types and number of mRNAs that a given sRNA can bind. Based on these factors Sharma et al.[7] (ref. 72 in Table 1) developed a high-throughput strategy for the engineering of synthetic sRNAs. In their approach, the Hfq domain was left unchanged and a library of randomized binding domains was generated. A natural 5’ UTR was fused to a reporter gene (GFP) and the researchers selected for the repression of this gene. They were able to successfully identify sRNA candidates that repress ompF and fliC mRNAs. Interestingly, the authors observed that the artificial constructs repressing the ompF exhibit important similarities in the features shown by the natural ompF repressor, the sRNA MicF (Figure 3). A recent work studied the free-energy of the complex sRNA-mRNA and found an important correlation between structure-function in sRNAs. Hao et al. [8] (ref. 104 in Table 1) generated numerous mutants of the sRNA RyhB and tested in vivo their gene control function. They concluded that when using a thermodynamic model to compute the free-energy of the mRNA-sRNA complex, these values exponentially correlated to the gene silencing strengths shown by the mutants.
sRNAs in metabolic engineering
Metabolic engineering is an enabling technology for strain optimization towards the production enhancement of biotechnological substances. As aforementioned, sRNAs are ideal candidates for developing and alternative methodology for the combinatorial knockdown of genes in metabolic engineering. Towards these purposes, Na et al.[9] (ref. 68 in Table 1) generated a library of artificial sRNAs that target a diversity of chromosomal gene targets. Then, by a combinatorial approach they isolated a strain that was able to substantially increase cadaverine production and tyrosine production. Specifically, the authors of this work selected the MicC sRNA scaffold, that includes the Hfq-binding site, and modify the binding domain by the introduction of anti-sequences of genes involved in the metabolic pathway of either cadaverine or tyrosine. Subsequently, they created a library of anti-sense RNAs and isolated the strains with higher production of the target molecules. Finally, used what they called forward engineering, to fine-tune the production optimization of these two metabolites by binding energy. They identified genes not expected to affect the titer of these metabolites but that are involved in the metabolic pathway regulation. This last realization represents a advantage over other traditional metabolic engineering approaches. In addition, this sRNA-based approach is generalizable to other bacterial strains. The strategies proposed by the authors possess important advantages over traditional gene knockouts methodologies due to the ability to fine-tune gene silencing, target multiple genes, easy-implementation and the ability to modulate gene expression without modifying those genes. These strategies avoid the burdensome generation of strain libraries.
As it can be confirmed from table 1, there are very few examples of the use of sRNAs for metabolic engineering applications. However, it is expected that this field will soon explode to produce numerous works and even applications aiming for more efficient strain optimization techniques for the production of biotechnologically relevant molecules.

A robust gene expression control device inspired on sRNAs

Isaacs et al.[10] developed a riboregulator system showing an enhanced dynamic range. This riboregulator design is inspired on the DsrA-RpoS sRNA system (Figure 4). This system has pioneered the field of rational design of sRNA-like systems and seeded a variety of applications based upon this same device e.g. a "cell that counts"[11] and a "switchboard"[12]. More recently, this cr-taRNA system has been used to test the influence of the Hfq assistance. Sakai et al.[6] introduced a Hfq domain into the taRNA and found improved results in gene expression control suggesting that in vivo Hfq enhances the inherent sRNA regulatory capacity.
Future directions for sRNAs in Synthetic Biology
To date, sRNA synthetic systems remain as a widely unexplored field moreover when referring to metabolic engineering applications. Examples of sRNA-inspired devices date back to 2004 and since then several artificial sRNA-like devices have been created, in its majority aiming for gene silencing applications. However, these pioneering examples, although claimed to have been inspired over natural sRNAs did not exploit in full sRNA features as sRNA were still very novel molecules. Recently, works such as the ones listed in Table 1 have been exploiting more deeply sRNA features for the gene silencing purposes. Definitely the work carried out by Na et al. [9] is a methodology for strain optimization with a great potential to be widely exploited in the metabolic engineering field. It is expected that this method will continue to be refined and standardized with the vision of using it in combination with traditional strain optimization techniques to enhance metabolic engineering ability to increase the production of relevant substances at the industrial scale. Although this work represents a great leap in the use of sRNA-based strategies in metabolic engineering, it did not exploit a very useful capability of sRNAs just yet: multi-targeting. In lieu of the recent interest in sRNA, it is plausible to expect that researches will start working on DsrA-like systems. DsRA is a sRNA that can control two target mRNAs at once as it activates production of RpoS mRNA (the stationary phase sigma factor) and inhibits H-NS (histone-like nucleoid-structuring protein) translation. This astonishing ability to repress and enhance the production of two different mRNAs a the same time seems of great relevance since for strain optimization some genes are turned on and some are turned down simultaneously for an overall increase in the production of the molecule of interest. To date, there are no examples of such an artificial sRNA with this dual capability. These promising perspectives at the same time are in the need of enabling technologies, the development of rational design approaches is of great relevance to assist on the sRNA rational design[4]. Finally, sRNAs have shown their potential use as metabolic target genes, as it can be confirmed from Na et al.[9] work, they were able to identify genes involved in the metabolic pathway of the metabolites of interest that were not expected to have an effect in the overall production. In addition, the fine-tuning capabilities of sRNA-like systems allows for the partial repression of essential genes without the negative consequence of inviable cells.
sRNA-like iGEM projects
The Denmark Technical University team in 2011 [13] used a bioinformatics approach to confirm the structural features present in an sRNA e.g. binding domain, Hfq domain, transcription terminator and linker region. They investigated the sRNA system chitobiose that requires the presence of another sRNA called trap-RNA (in this case chiXR) to release the silencing imparted by chiX on its target mRNA chiP. This work represents an interesting confirmation experiment of what had been already reported in the literature since they inserted chiP in a plasmid a showed that its expression was regulated by chiX and when changing the complementary binding region the regulation is removed.
Other teams such as the Ocean University of China iGEM 2012 [14] team aimed to develop a decision-making device based on sRNA regulation to predict when red tide is going to happen. In another example, Uppsala University iGEM 2012 team [15] constructed synthetic sRNAs that can down regulated antibiotic resistance genes by engineering the binding domain of the sRNA Spot42.
References
Error fetching PMID 20980440:
Error fetching PMID 21925377:
Error fetching PMID 23362267:
Error fetching PMID 23651005:
Error fetching PMID 21742981:
Error fetching PMID 23334451:
Error fetching PMID 24356572:
Error fetching PMID 15208640:
Error fetching PMID 24328142:
Error fetching PMID 22454498:
Error fetching PMID 19478183:
- Error fetching PMID 15487940:
Comprehensive review on bacterial small RNAs - Error fetching PMID 20980440:
A more recent review on bacterial small RNAs. - Error fetching PMID 21925377:
Another recent review on bacterial small RNAs. - Error fetching PMID 24356572:
A thorough review on synthetic regulatory RNAs. - Error fetching PMID 23362267:
A review on sRNA negative regulation. - Error fetching PMID 24328142:
Effect of Hfq domain introduction into a synthetic sRNA. - Error fetching PMID 23651005:
High-throughput method for the engineering of sRNAs. - Error fetching PMID 21742981:
sRNA structure-function relationship. - Error fetching PMID 23334451:
sRNAs in metabolic engineering. - Error fetching PMID 15208640:
A robust sRNA-inspired riboregulator. - Error fetching PMID 19478183:
A transcriptional cascade based of an sRNA-like device that counts up to three. - Error fetching PMID 22454498:
A genetic switchboard based on an sRNA-like device. - [http://2011.igem.org/Team:DTU-Denmark/Project
sRNA system with a trap-RNA for chitibiose control. - [http://2012.igem.org/Team:OUC-China/Project/Overview
sRNA system for the prediction of red tide. - [http://2012.igem.org/Team:Uppsala_University
sRNA system for the repression of resistance genes in bacteria.