CH391L/S14/ApplicationsofOptogenetics
Contents |
Introduction to Optogenetics
Optogenetics is a field of study that employs techniques from optics and genetics to control and study the functions of cells. In short, optogenetics involves the integration of extracellular genes to develop light sensitive proteins, strategies for targeted illumination, and finally, controlled readouts for reporting on changes in the cell. [1]
A brief understanding of how optogenetics is applied; the 4 general steps for creating a light sensitive system:
- Step 1: Identify potential light sensitive proteins: Light sensitive proteins can be naturally occurring or chemically modified to become photosensitive. Photosensitive light proteins can be used to modulate membrane potential or modulate cell signaling. Examples will be provided later.
- Step 2: Delivering the genes: Genes coding these light sensitive proteins can be delivered to the target cells by transfection (deliberately inserting nucleic acids via non-viral techniques such as electroporation),[2] viral transduction, or creation of transgenic animal lines.
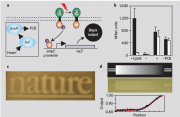
- Step 3: Controlled illumination: Control of cellular activity depends on the well-defined temporal and spacial control of the illumination of light. There are several techniques for different desired effects. The 2012 Washington iGEM team designed a particularly useful application called "E. colight" that can control the illumination delivered to both a petri dish and a 96 well plate. [3]
- Step 4: Reading the outcome: The effect induced by illumination of the photosensitive proteins must be measured in cells, tissues, and organisms. For example, electrodes can be used to monitor effects of membrane voltage.
The basics of optogenetics can be found here, at a previously created page from 2013.
Applications
Engineering E. Coli To See Light
Levskaya et al reported in 2005 a bacterial system in Nature Magazine where they created a bacterial system that is dependent on red light to switch states.
How was this system made? Using light to modulate cells is not a new concept. For millions of years, plants have interacted with light using phytochromes which are photoreceptive signaling proteins responsible for mediating seed germination, seedling de-etiolation, shade avoidance, and photosynthesis. However, E. Coli and other enterobacteria do not have photoreceptors, so for this experiment a chimaera was created that uses a phytochrome from a cyanobacterium. A phytochrome is a two-component system that is made up of a membrane bound, extracellular sensor that responds to light, and an intracellular response regulator. In this case, the response regulator was bound to an E. coli intracellular histidine kinase domain. The next challenge faced was that the part of the photoreceptor that responds to light, phycocyanobilin, is not naturally produced in E. Coli. So, two phycocynaobilin-biosynthesis genes were inserted, and a pathway was introduced that can convert haem into phycocyanobilin. The working phytochrome modulates the activity of the ompC promoter, thus controlling the gene it promotes: the lacZ gene. [4] The synthetic sensor kinase phosphorylates the ompC promoter which allows the lacZ to be expressed. When exposed to light, the kinase is inhibited from phosphorylating the ompC promoter, and thus the lacZ gene cannot function. This is exemplified by the picture to the right of the "Nature" text. The bacteria were placed on an agar plate that contained S-gal. When functional, lacZ catalyzes the formation of a stable, insoluble, black precipitate from S-gal. The part in the middle was exposed to red light, thus inhibiting the bacterias ability to express the lacZ gene, which is why it is white. The surrounding black was not exposed to light, so the lacZ gene could function and degrade the S-gal.
Using light to modulate cells is not a new concept. For millions of years, plants have interacted with light using phytochromes which are photoreceptive signaling proteins responsible for mediating seed germination, seedling de-etiolation, shade avoidance, and photosynthesis. However, E. Coli and other enterobacteria do not have photoreceptors, so for this experiment a chimaera was created that uses a phytochrome from a cyanobacterium.
A phytochrome is a two-component system that is made up of a membrane bound, extracellular sensor that responds to light, and an intracellular response regulator. In this case, the response regulator was bound to an E. coli intracellular histidine kinase domain. The next challenge faced was that the part of the photoreceptor that responds to light, phycocyanobilin, is not naturally produced in E. Coli. So, two phycocynaobilin-biosynthesis genes were inserted, and a pathway was introduced that can convert haem into phycocyanobilin. As described above, the working phytochrome modulates the activity of the ompC promoter, thus controlling the lacZ gene expression. [4]
Neuroscience
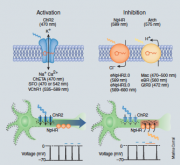
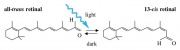
Optogenetics can be a powerful tool for monitoring the activity of individual neurons in living tissue. It uses light to control the activity of neurons that have been modified to express light sensitive proteins. There are two important light-sensitive proteins that modulate the activity of neurons: channelrhodopsins, and halorhodopsins, [5] that are both part of the opsin family of proteins. Opsins are activated by the absorption of a photon of light that results in the photoisomerisation of the chromophore from the -cis to an all-trans conformation. The photoisomerization induces a conformational change in the opsin protein, causing the activation of the phototransduction cascade.
- Channelrhodopsins: are cation channels that produce depolarization of neurons when illuminated. They function as light gated ion channels. Channelrhodopsins originally served as sensory photoreceptors in unicellular algae. The channelrhodopsin that is utilized in optogenetics was isolated from Chlamydomonas reinhardtii. [6]
- Halorhodopsins: are another light gated ion channel (that pumps chlorine) that can be used to inhibit the action potentials of mammalian neurons. Pumping chlorine into the cells hyperpolarizes it, thus inhibiting action potentials.[7] Halorhodopsins are found in phylogenetically ancient archaea, known as halobacteria. The halorhodospin that is utilized in optogenetics was isolated from the halophilic bacterium Natronomonas pharaonis.
Optogenetics is advantageous because other neuromodulation techniques either simultaneously affect surrounding cells and processes, or have slow kinetics and reversibility. Microbial opsins have fast on-off kinetics, which makes it possible to evoke or inhibit neural activities within milliseconds (which is a time scale relevant to that of the brains typical physiological functions). Additionally, selective expression of the opsins in targeted neurons allows for specificity. These two advantages make optogenetics the technique of choice when studying neurons and brain function. [8] [9]
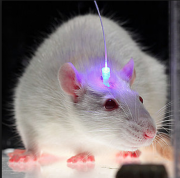
Why Mice? Mice are an excellent mammalian organism to do biomedical research. The physiology shared between mice and humans makes the mouse ideal for modeling complex human diseases and testing optogenetic systems.
The optogenetic process:
1. Opsin gene is combined with a promotor to ensure the gene is active only in a specific type of cell 2. The modified gene is inserted into the host brain via a viral vector 3. The virus is nonspecific when infecting cells, but the promoter only allows selected neurons to make opsin protein. 4. Fiber optic or optrode is inserted into the brain and flashes a specific wavelength of light 5. Record transformed regions.
How do mice become optogenetic test subjects? This brief video illustrates the process of outfitting the mouse with the fiber optic cable and genes necessary for control.
Optogenetic control of epileptiform activity
Epilepsy is a neurological condition that is characterized by recurrent seizures that affects nearly 1% of the general population. Current treatments have limited success and high direct and indirect costs of treatment. In this experiment, a halorhodopsin chloride pump (NpHR) was proposed as a solution for epilepsy . Genetically adapted to be expressed in mammalian brain by viral vectors and optically activated with millisecond precision, the transgene NpHR pumps chloride ions into transduced neurons, hyperpolarizing and preventing them from firing action potentials. Being able to control the excitability of a cell could be an powerful tool to prevent excessive activation of neurons in pathologic conditions within specific brain regions (e.g., an epileptic seizure focus). Jan Tonnesen et. al. writes:
"We used a lentiviral (LV) vector containing the NpHR gene driven by the calcium/calmodulin dependent protein kinase II-alpha (CaMKII-alpha) promoter, which expresses the transgene selectively in the principle glutamatergic neurons of the hippocampal formation."
They found that the NpHR transduced neurons were hyperpolarized and silenced by exposure to orange light of 573-613-nm wave-length that activated the transgene NpHR. Jan Tonnesen et. al. proposes that this could be considered a reliable model for testing different treatment approaches for suppressing pharmacoresistent epilepsy as well as potentially being a solution for patients with temporal lobe epilepsy. It is reasonable to speculate that implantable optogenetic devices can be coupled to implantable seizure onset-predicting devices that would trigger optical stimulation in a timely manner to prevent forthcoming epileptic episodes. [10]
Future Directions
In 2010, a collaborative project called OpenOptogenetics was created by Guillaume P. Dugué, in an effort to promote, facilitate, and democratize the use of optogenetic approaches in biological research. The website states on its homepage:
"OpenOptogenetics provides background knowledge, an inventory of available optogenetic tools and their characteristics, tips and protocols, reviews of commercially available equipment and bibliographic references. OpenOptogenetics is an open wiki, just like Wikipedia, and therefore is also an attempt to promote “open research”, where researchers make clear accounts of their methodology and share them with their colleagues without the intermediary of a publication."
OpenOptogenetics could be a powerful tool when working with optogenetics in future research. [11]
iGEM applications
As stated before, the 2012 Washington iGEM team used optogenetics for protein regulation that functioned similarly to the Levskya et. al. experiment. They also developed techniques that allowed them to use technology to provide the light to modulate the cells. They cleverly called their app "E. colight.". Their overall project for that year was named "Apptogenetics," as they had developed many different types of technological apps to aid them in their experiments. [3]
The University of Technology in Munich Team in 2010 created a bacterial constructed 3D structures by using light induced gene expression. This experiment also integrated non-canonical amino acids and recoding of the amber codon, and it could be useful for the 2014 iGEM team to look into their research. [12]
References
- Pastrana, E. "Optogenetics: controlling cell function with light." Nature Methods 8, 24–49 (2010).
Basics of Optogenetics - Promega http://www.promega.com/resources/product-guides-and-selectors/protocols-and-applications-guide/transfection/
Transfection information - iGEM Washington team http://2012.igem.org/Team:Washington/Optogenetics
E. Colight/Apptogenetics - Levskaya, A. et al. "Synthetic biology: engineering Escherichia coli to see light." Nature 438, 441–442 (2005).
Engineering E.Coli to see light - Lalumiere, R. "BRAIN STIMULATION: Basic, Translational, and Clinical Research in Neuromodulation" - January 2011 (Vol. 4, Issue 1, Pages 1-6, DOI: 10.1016/j.brs.2010.09.009)
Channelrhodopsins and Halorhodopsins - OpenOptogenetics http://www.openoptogenetics.org/index.php?title=Main_Page
OpenOptogenetics homepage - Yasuo Mukohata, Kunio Ihara, Takeshi Tamura, and Yasuo Sugiyama "Halobacterial Rhodopsins" J Biochem (1999) 125 (4): 649-657
Halorhodopsin information - Fenno, L., Yizhar, O., Deisseroth, K. (2011) "The Development and Application of Optogenetics" Annu. Rev. Neurosci. 34, 389-412
General background information over tools and techniques - JY. Mei, F. Zhang "Molecular tools and approaches for optogenetics" Biol. Psychiatry, 71 (2012), pp. 1033–1038
Advantages of optogenetics - J. Tønnesen, A.T. Sørensen, K. Deisseroth, C. Lundberg, M. Kokaia "Optogenetic control of epileptiform activity" Proc. Natl. Acad. Sci. USA, 106 (2009), pp. 12162–12167
Optogenetic control of epileptiform activity - Munich 2011 iGEM team http://2011.igem.org/Team:TU_Munich/project/introduction
Printing and light sensors