CH391L/S14/Artemisinin
Contents |
Artemisinin
Artemisinin is a natural plant product whose derivatives form the basis for antimalarial medication.History
Also known as wormwood or quinghao, artemisinin was used in ancient China to treat fevers and what is now known as malaria. The compound was rediscovered in 1970s as part of an effort to fight strains of malaria that had become resistant to contemporary medications. Artemisinin derivatives with improved pharmacokinetic properties were developed, and by 2002 the World Health Organization had named these compounds the first line in defense against uncomplicated malaria caused by Plasmodium falciparum. [1]
Artemisinin is traditionally isolated in very low yields from the plant Artemesia annua. Additionally, approximately 18 months pass between planting and isolation of artemisinin. Fluctuations in supply and high costs threaten the availability of the compound in the developing countries where it is most needed.[1] Artemisinin therefore represents an important synthetic target. A successful synthesis would produce artemsinin, or a precursor, on the order of 25 g/L to be able to supplement global supply in a meaningful and feasible way. [1] The most successful strategy thus far, outlined below, has been to engineer microbes to produce an intermediate on the path to artemisinin which is converted by chemical reactions into artemisnin.
Semisynthetic Route
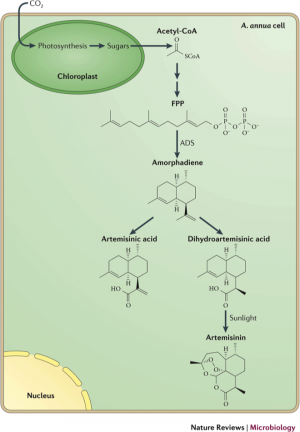
Optimization of amorphadiene production
The first committed intermediate in the route to artemesinin is amorphadiene, which is derived from the isoprenoid farnesyl diphosphate (FPP). To increase overall flux into the artemisinin synthesis pathway, enzymes of the 1-deoxy-D-xylulose 5-phosphate (DXP) pathway, which produces FPP in E. coli, were overexpressed in that organism. This strategy increased production of pathway intermediates, but only to low mg/L amounts. [2]
In yeast, FPP is produced from an alternate route- the mevalonate pathway. Martin and coworkers [3] split the genes encoding the mevalonate pathway onto two plasmids (MevT and MevB) and expressed them in E. coli, as shown on the right side of Figure 3, below. They also expressed an ADS enzyme optimized for E. coli, which as shown in Figure 2 converts FPP to the amorphadiene intermediate. Strains expressing the mevalonate plasmids and the ADS, with optimized fermentation conditions, produced 0.5 g/L amorphadiene.
Studies by Pitera and coworkers showed that increased expression of the mevT operon caused inhibition of growth.[4] More specifically, accumulation of an intermediate in the part of the mevalonate path that was encoded by MevT plasmid inhibited fatty acid biosynthesis. To address this issue, the expression of enzymes just upstream and downstream from the intermediate was reduced and increased, respectively. Along with that change, replacement of the downstream enzymes with genes from other species resulted in production of amorphadiene at concentrations of about 25g/L . [5] [6]
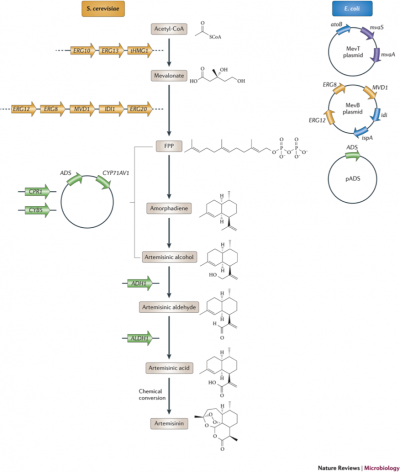
Optimization of artemisinic acid production
Once acceptable amorphadiene production levels were achieved, efforts were made to increase production of the more direct precursor artemisinic acid. In A. annua, amorphadiene is converted to more oxidized compounds, including artemisinic acid, by a cytochrome P450 (CYP71AV1). When expressed in amorphadiene-producing S. cerevisae, CYP71AV1 was shown to accomplish the conversion.[7] However, E. coli is generally unable to express eukaryotic P450 enzymes. Parallel tests of E. coli and S. cerevisae found that S. cerevisae produced higher concentrations of artemisinic acid (2.5 g/L vs 1 g/L for E. coli) under conditions suitable for scaling [8] and was therefore chosen as the host organism.
Once yeast was determined to be the best organism, a strain with desirable properties (CEN.PK2) was chosen and its mevalonate pathway was overexpressed to increase amorphadiene production. Further process improvements led to a strain producing 40 g/L of amorphadiene. [9] However, expression of CYP71AV1 did not lead to increased artemisinic acid levels.
Cells that expressed CYP71AV1 were found to exhibit decreased viability. The possibility that poor interaction between CYP71AV1 and its cognate reductase was creating destructive reactive oxygen species merited a decrease in the expressed reductase. [1] [10] Additionally, cytochrome b5 from A. annua was expressed to increase the rate of P450.[11] These changes were beneficial in amount of artemisinic acid but produced high levels of the presumably toxic intermediate artemisinic aldehyde. [10]
Drastic improvement in the production of artemisinic acid was achieved through coexpression of artemisinic aldehyde dehydrogenase and artemisinic alcohol dehydrogenase, both from A. annua. When the mevalonate pathway was constitutively expressed in this strain, Paddon et al report that the target concentration of 25 g/L arteminisic acid was achieved. [10]
Conversion of artemisinic acid to artemisinin
The route from artemisinic acid to artemisinin is shown in Figure 4.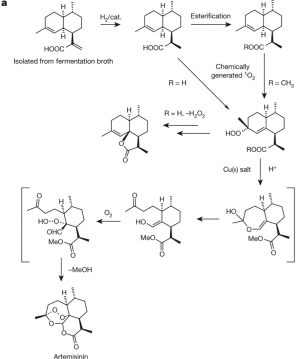
Future directions
This method of utilizing reaction pathways in nature to achieve complex chemical transformations only becomes more exciting as new synthetic biology tools are developed. More efficient ways to study and manipulate genetic systems will allow access to much more difficult and complicated structures. Paddon [1] illustrates other molecules whose synthesis might be aided by a similar semisynthetic route, including vincristine, morphine, and prostratin. The artemisinin semisynthesis was successful in part due to thorough understanding of the pathway used. Targets with well known biosynthesis pathways may be more likely to succeed.
Other strategies
While the semisynthetic route has been most successful, there are other strategies. For example, Zhu et al completed a total synthesis of artemisinin.[12] While their starting material was inexpensive, it is not ready to be scaled up to global supply. Others have attempted to use transgenic approaches to improve artemisinin yields in A. annua or other plants, such as tobacco. These efforts seem to mimic the same strategies embraced the semisynthetic Keasling method in a plant host (ie overexpress key enzymes). While improvements have been made, they are not on the order of the semisynthetic route. [13]
iGem
VCU (2011) attempted to introduce isoprenoid pathways into Synechococcus elongatus for possible use as host organisms. Success in this endeavor was not clear.
References
Error fetching PMID 15236249:
Error fetching PMID 12778056:
Error fetching PMID 17239639:
Error fetching PMID 16845378:
Error fetching PMID 19221601:
Error fetching PMID 16612385:
Error fetching PMID 19194910:
Error fetching PMID 22247290:
Error fetching PMID 23575629:
Error fetching PMID 17693640:
Error fetching PMID 22866604:
Error fetching PMID 24413765:
- Error fetching PMID 24686413:
Excellent review of the story of the successful artemisinin semi-synthesis from Keasling. - Error fetching PMID 15236249:
Slight increase in terpenes in E. coli through overexpression of DXP pathway. - Error fetching PMID 12778056:
Introduction of mevalonate pathway and optimized ADS to E. coli. - Error fetching PMID 17239639:
Identified mevT operon expression as inhibiting E.coli growth. - Error fetching PMID 16845378:
Changed enzyme expression levels to reduce HMG-coA accumulation. - Error fetching PMID 19221601:
Achieved amorphadiene concentration of 25 g/L. - Error fetching PMID 16612385:
Isolated CYP71AV1 as amorphadiene oxidizer in A. annua; demonstrated its function in S. cerevisae. - Error fetching PMID 19194910:
Determined S. cerevisae was a better organism for industrial artemisinic acid production. - Error fetching PMID 22247290:
Achieved production of amorphadiene at 40 g/L by strain engineering and other measures. - Error fetching PMID 23575629:
Achieved production of artemisinic acid at 25 g/L and converted to artemisinin with good yield. - Error fetching PMID 17693640:
Describes role of cytochrome b5 as increasing the rate of cytochrome P450. - Error fetching PMID 22866604:
Total organic synthesis of artemisinin. - Error fetching PMID 24413765:
Describes efforts to increase plant-based synthesis of artemisinin.