CH391L/S14/Biocontainment
Contents |
Biocontainment
Biocontainment describes efforts to keep separated laboratory synthesized- and naturally synthesized- genetic information. While physical containment (appropriate bioreactor design, safety education and training of scientists/workers) of laboratory samples is a necessary aspect of biocontainment, this article will focus on efforts to create chemical barriers between laboratory-synthesized and naturally occuring genetic information sources.
History and Context
While recombinant DNA technology in 2014 is widespread, during its early stages in 1975 scientists were facing a scenario very similar to today. The emerging use of restriction endonucleases then represented manipulation of DNA without knowing the consequences of possible release into the environment. The Asilomar Conference, called to address these worries, recommended very cautious use of the new science. The legacy of the Asilomar Conference includes the Biosafety level designation as well as an abundance of evidence that recombinant DNA experiments in general pose little to no risk in the case of escape. [1] Keeping in mind the fundamental differences between the emerging science of 1975 and today, the Asilomar conference could provide a blueprint for how to consider safety implications of new technologies.
Active Containment Strategies
Endeavors to isolate laboratory synthesized genetic information fall into two broad classes: 1) genetically encoded dependence systems and 2) semantic firewalls.
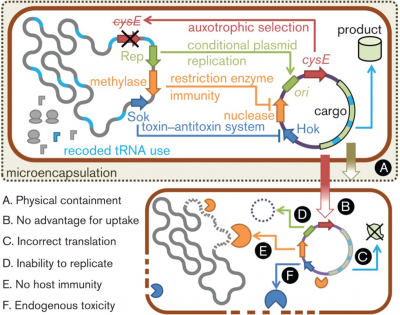
Dependence Systems
Dependence systems seek to use translation of the synthesized organism against itself. Elements are introduced into the synthesized genetic material so that it will not be able to survive outside its designed host cell.
- Toxin/Anti-toxin Pairs:[2]
- This strategy represses toxin production until death is desired. For example, a toxin gene located downstream of a repressible promoter that may be induced upon addition of an certain molecule. Another variety of this strategy would express the toxin and instead control the expression of the anti-toxin. If the toxin were plasmid-encoded, it would poison an organism that acquired it by HGT. These strategies are susceptible to random mutations that make the toxin inactive.
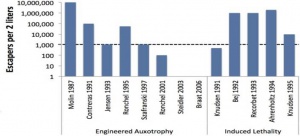
- Engineered Auxotrophy:
- Designing auxotrophy into genetic material forces it to be dependent on some laboratory supplied molecule (for example, an amino acid). As the entire gene is missing, random mutations will not inactivate the system (ie supply the metabolite) as above. However, in the case of release in the environment, auxotrophic organisms may be able to scavenge the necessary ingredients.
- Amber Suppression:
- A variation on auxotrophy, this system places an amber termination codon upstream of the gene to be knocked out. Supplementation with a novel aminoacyl-tRNA synthetase-tRNA pair allows read-through past the amber codon resulting in no auxotrophy. If an organism with this system were to escape or be released, the absence of the aminoacyl-tRNA synthetase-tRNA pair would result in truncation. Pairing the complementary codon to a non-canonical amino acid further separates the synthetic organism from natural organisms. Inefficient amber suppression, mutation of the amber codon and unintended effects on other amber codons in the genome are drawbacks to this method.[3]
Semantic Firewalls
The goal of a semantic firewall is to completely isolate synthetic organisms from nature through the use of novel genetic information systems. The structure of DNA can be changed in a number of ways to make it incompatible with the largely conserved natural replication machinery. These new molecules would be meaningless if taken up by natural organisms and would likely be ejected.[1]
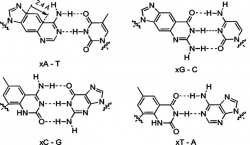
- Base Pair Size Change
- Homologating the base pairs (ie, add an extra benzene ring) may to render them unfit to be used by the existing replication machinery. The hydrogen bond pattern of the altered bases is unchanged (some work has changed that by using other tautomers of the bases), allowing the new bases to pair with natural ones, but altered base would preclude replication with the natural polymerases, which have evolved to DNA. This method would require development of specific polymerases to fit the modified base pairs. It would also be necessary that the new molecules do not tolerate the natural DNA polymerases well. For example, Krueger et al. have described selection of a polymerase that is functional with homologated base pairs. [7]
- Codon Reassignment
- Another strategy would be to render certain nonsense codons redundant, then introduce ways for those codons to code for unnatural amino acids. This idea was employed by Isaacs et al. to replace all TAG stop codons with a redundant TAA to free the TAG to code for a non-natural amino acid. This work shows that organisms can successfully be significantly redesigned for our purposes. With the proper synthetases, these codons could be powerful tools for designing proteins but would not be translated by outside machinery. [8]
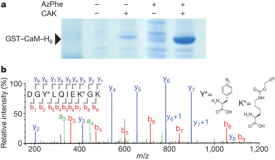
- Quadruplet Codons
- If a novel ribosome was designed to accommodate four base-pair codons, translation of mRNA would give a vastly different output than a canonical three base-pair codon. In this instance, both the ribosome and the mRNA would be orthogonal to the “natural” organism. In addition, a four base-pair codon would create many more recognition sequences, enabling further differentiation from current proteins through the association of new amino acids with the open codons. Neumann and coworkers successfully selected for a ribosome that could recognize four base pair codons. Additionally, they were able to synthesize a protein incorporating unnatural amino acids that would only be fully translated if those amino acids were supplied.[9]
While promising in theory, we are still far from bringing these semantic firewalls into place. Much work needs to be done to understand and synthesize the machinery required for these alternate genetic sequences. Given that interaction with the environment can be expected, it is important to consider at each step the capabilities of the molecules synthesized in the lab.
iGEM
In recent years, iGEM teams have developed a multifaceted physical, dependent, and semantic system (Paris Bettencourt 2012), a toxin/anti-toxin system (UCSF 2012), and developed GeneGuard to minimize HGT (Imperial College 2011).
References
Error fetching PMID 23519158:
Error fetching PMID 20307192:
Error fetching PMID 11375176:
Error fetching PMID 3554958:
Error fetching PMID 21981660:
Error fetching PMID 21764749:
Error fetching PMID 20154731:
- Error fetching PMID 22710182:
Review of biocontainment strategies, especially orthogonal systems - Error fetching PMID 23519158:
Another review of biocontainment. More information about dependent systems and a generally wider scope. - Error fetching PMID 20307192:
Describes advances in systems that incorporate unnatural amino acids - Error fetching PMID 3554958:
Review. Special focus on the role synthetic biology plays/should play in wider society. - Error fetching PMID 11375176:
Modified an organism for to have two routes of dependence on 3-methylbenzoate - Error fetching PMID 21981660:
Change base to include internal benzene ring, resulting in wider base pair - Error fetching PMID 21764749:
Changed all TAG codons in various E. coli strains to TAA - Error fetching PMID 20154731:
Selection of a ribosome that can read four base pair codons