CH391L/S14/NitrogenFixationRefactoring
Contents |
What is Gene Refactoring?
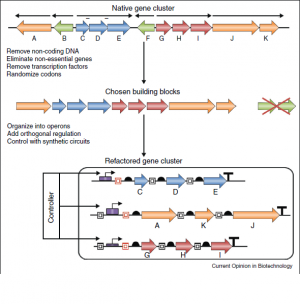
Prokaryotic genes associated with a specific function are often grouped together in contiguous regions of the genome known as gene clusters [1]. Many of the functions encoded in these clusters are of potential interest to synthetic biology. Unfortunately, gene clusters are frequently subject to complex and highly redundant host regulation [1]. Furthermore, there may be no laboratory conditions known that allow for expression of these clusters. However, through a process known as “refactoring,” gene clusters can be recoded to systematically eliminate native regulation [2]. This process of refactoring aims to reduce the overall complexity of genetic systems and allows them to be tailored for a particular purpose. The term “refactoring” is borrowed from computer science and refers to the alteration of a program’s underlying code without changing its functionality [2]. This term was first applied in biology to describe the top-down approach of simplifying the phage T7 genome [3], but it is used here to refer to the bottom-up approach of eliminating native regulation of gene clusters and "replac[ing] it with synthetic genetic parts and circuits" [2]. The result of this process is is a simplified genetic system whose DNA sequence has been altered but encodes for the same function [2]. Operon refactoring is essentially a more systematic approach to metabolic engineering that aims to generate a simplified operon free of any internal and often cryptic regulation.
Process
Find A High Priority Gene Cluster
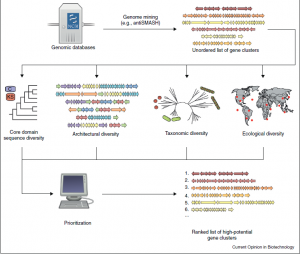
One of the biggest challenges that is faced with this area of study is to search for a high-priority gene cluster. This proves to be difficult because many organisms, containing certain attractive secondary metabolic pathways, are impossible to grow in a laboratory setting. One way to circumvent this problem is to develop a high-throughput method of searching for gene clusters across the tree of life. If a mathematical algorithm could be created to search for high-priority gene clusters, novel chemical pathways with industrial potential could be found. However, this proves to be incredibly difficult. Instead, many refactored gene clusters come from well-characterized chemical cascades that have industrial or medical potential. One of these pathways include the nixtrogen fixation gene cluster.
Select A Suitable Host Organism
The next step in gene refactoring is to find a suitable host that would readily overexpress the desired gene products. There are several criteria that an organism must be meet in order to be considered a suitable host. A host must be easy to manipulate, it must offer a clean background, and it must be easily cultured. Current available host strains (e.g E.Coli, streptomyces, bacillus, etc) are good starting host organisms, but they are not necessarily optimized for the specific refactored gene cluster. There are several strategies available to find an optimized host to express a certain cluster. One of these strategies is to find a host cell that already produces a similar product. However, even if an optimal host organism is found, a large amount of additional engineering is required for optimization. The potential host organisms must be able to grow readily on cheap nutrients, grow at high densities, and must be easy to manipulate.
Remove All Native Regulation
After a suitable host organism is selected, the gene cluster must be "refactored" in order to both optimize heterologous expression and to remove all native regulation. A gene is refactored to optimize expression by using codons that are commonly used in the prospective host organism. Another way to optimize heterologous expression is to replace its native start codon with a start codon that is more commonly recognized in the new host organism. Much of the gene functions will remain the same in a refactored gene cluster, but codon usage and overall architecture of the genetic code will be largely altered. The purpose of altering codons is to disrupt any native regulatory sequences that are hidden within the genetic code. These regulatory sequences include regulatory genes, promoters, ribosome binding sequences, small RNAs, and RNA secondary structures. Once the codons were computationally changed, a computer was used to scan the sequence to ensure that a known regulatory sequence wasn't accidentally introduced into the refactored gene cluster.
Place Cluster Under Synthetic Control
Once the native regulatory sequences were removed, the genes were placed under synthetic control. New inducible regulatory circuits, which are orthogonal to native regulation of the host, must be introduced in order for the genes to be expressed. For a synthetic pathway to function correctly, the operons must be expressed in the correct stoichiometry. This can be accomplished by introducing promoters with different strengths to express correct levels of gene product. Another way that the stoichiometry of the refactored gene cluster could be manipulated is through introducing ribosome binding sequences of different strengths. In this schematic, the same operon will be used to transcribe the gene, but different ribosome binding sequences will be used at every gene to translate the correct amount of product.
Assemble The Parts
The theorized refactored gene cluster must then be assembled in order to be expressed in host organisms. There are special challenges that must be overcome to assemble such large chunks of DNA. The enzyme complexes in many interesting secondary metabolite pathways include over 10,000 amino acids and are highly repetitive due to their multimodular structure. It is unlikely, therefore, that these large sequences could be assembled in vivo using yeast because many of the repetitive sequences lead to deletion of parts by recombination events.
Nitrogen Fixation Refactoring
Tenme paper chose to refactor nitrogen fixation gene cluster of klebsiella oxytoca. Nitrogen fixation is the conversion of N2 into ammonia and is very industrially relevant because fertilizers are made from ammonia. K. oxytoca's gene cluster is an optimal cluster to refactor because it has been a model system for studying biological nitrogen fixation. It consists of 20 genes in 7 operons in 23.5 kb of DNA. Nitrogen fixation is tightly regulated by fixed ammonia, oxygen, and temperature. Therefore, it was important to remove all native regulatory sequences before being able to be heterologously expressed in E. coli. Tenme et al did a robustness test to see what kind of promoters were needed for optimal stoichiometry. The group knocked out genes in the WT background and complemented them under inducible control. Using different promoter strengths to express the complemented genes, clear optimums were able to be produced. From the data, the group decided to place the genes into four operons with each gene having a RBS and insulator sequences to separate genes. The group decided that some genes were not necessary for nitrogenase activity to occur, so they left them out (nifT, nifX, nifLA). An acetylene reduction assay was used to measure the nitrogenase activity of the refactored gene cluster. Only 7.4% of WT nitrogenase activity was recovered. The host E. coli also grew 3.5 times slow than the wild-type. However, reduced activity was expected when trying to simplify and modularize highly evolved systems. Although the refactored gene cluster did not produce a large quantity of nitrogenase activity, the refactored genes can be used for studying basic biological principles. Studying these refactored genes can quantitatively show how adding back certain regulation steps can increase nitrogenase activity by starting with a clean reference point (a cluster that removed all native regulation). It can also be used to study the importance of temporal control or the need for genes to arranged with a particular operon structure. Finally, studying these refactored gene clusters can be used to discover novel genetics and regulatory modes.
- Introduction of pathway into cereal crops
iGEM Connection
The 2012 iGEM team at the University of Texas at Austin refactored the alkylxanthine degradation (Alx) gene cluster from Pseudomonas putida CBB5 to function as a caffeine-degrading operon in E. coli. E. coli strains harboring this operon were able to degrade caffeine to the guanine precursor, xanthine. Cells lacking a de novo guanine biosynthetic pathway and complemented with the refactored operon required caffeine for growth. Thus, these cells were shown to be “addicted” to caffeine [6].
References
- Frasch HJ, Medema MH, Takano E, and Breitling R. Design-based re-engineering of biosynthetic gene clusters: plug-and-play in practice. Current Opinions in biotechnology, 2013.
- Fischbach M and Voigt CA. Prokaryotic gene clusters: a rich toolbox for synthetic biology. Biotechnol J, 2010.
- Temme K, Zhao D, and Voight CA. Refactoring nitrogen fixation gene clusters from Klebsiella oxytoca. Proc Natl Acad Sci U S A, 2012.
- Chan LY, Kosuri, S, Endy D. Refactoring bacteriophage T7. Mol Syst Biol, 2004.
- Gibson DG, Young L, Chuang R, Venter JC, Hutchison CA, Smith HO. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat Methods, 2009.
- Quandt EM, Hammerling MJ, Summers RM, Otoupal PB, Slater B, Alnhhas RN, Dasgupta A, Bachman JL, Subramanian MV, Barrick JE. Decaffeination and measurement of caffeine content by addicted Escherichia coli with refactored N-demethylation operon from Pseudomonas putida CBB5. ACS Synth Biol, 2013.