CH391L/S14/BiologicalpartsandtheiGEMregistry
Contents |
Introduction
Biological parts are refined, functional DNA sequences that have been standardized so as to act the same way in any biological device. These DNA sequences may be coding regions from genes, regulatory elements, or other genetic elements.[1] The Registry of Standard Biological Parts, or iGEM Registry, is a collection of these parts, commonly referred to as BioBricks™, that is being expanded constantly and can be used freely to create biological devices. Located at the Massachusetts Institute of Technology, the registry stocks BioBricks and biological devices that have been previously designed and accepts any new BioBricks (that adhere to the specifications) that are submitted.
History
With the development of techniques and other advancements in the manipulation of DNA and RNA both in living organisms and out, genetic engineering has grown rapidly since the 1970s. Biological devices, DNA structures synthesized from various natural sources with a set function that draws on all of its parts, started to emerge but were not very cohesive, which led to difficulties in combining them. In 2003, Tom Knight, a researcher at MIT, developed the idea of the BioBrick, a standardized design for biological parts that can be combined with each other with no complications or unforeseen results.[2] Knight's idea was to apply the principles of mechanical design to genetic engineering – that is, making it so that each part is easily interchangeable without losing functionality. Around the same time at MIT, the International Genetically Engineered Machine (iGEM) competition began, allowing these iGEM teams to create biological devices and compete to see who's device was the best, giving incentive to create these biological parts. As the competition increases each year, the parts and devices from the previous years are stored and available for use, and the number and quality of parts increases. The parts are stored in the Registry of Standard Biological Parts using the BioBrick format, and can be ordered by iGEM teams or synthetic biology labs.
Standardization
Standardization is key to the iGEM registry's system. Without the standards set by the registry, the BioBricks would not be able to be combined in different ways and still function the same. The key to standardization for BioBricks is the way each one is assembled: there is a very specific set of parts in each BioBrick which allows them to be combined easily and in any order.[2] An important idea for standardizing the process of making BioBricks is idempotence. Idempotence when used in terms of synthetic biology means that “each reaction leaves the key structural elements of the component the same.”[2] In other words, the BioBricks can be exposed to most reactions and environments without changing its key structure and function. This makes the parts much more reliable and easier to work with. Related to the idea of standardization is abstraction, another key feature of synthetic biology and the iGEM registry.[3] Abstraction refers to the idea of taking things out of context and looking at just the core features and behaviors. With regards to synthetic biology, that means that while the function of each individual DNA sequence may seem very complex, there are ways to categorize DNA sequences to allow for smoother usage and classification of biological parts. By grouping parts and devices on different levels of complexity, ranging from the overall category they belong to (such as promoter or terminator) to the very specific function that they perform, new parts and devices can be categorized more quickly and determining which parts could be interchanged in devices becomes much easier.
BioBricks
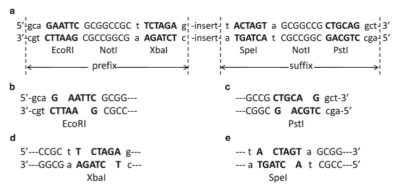
BioBricks consist of a few set components, plus the specific functional DNA sequence that will be used in other biological devices. Each BioBrick must have a set prefix and suffix: the prefix contains EcoRI, NotI, and XbaI restriction enzyme sites, while the suffix contains SpeI, NotI, and PstI restriction enzyme site. Because the sticky ends left by the XbaI and SpeI restriction sites are complementary, they can bind to each other to form a mixed site where neither restriction enzyme can bind.[4] This means every BioBrick has a common point to bind to the next that will not be broken if exposed to either of the restriction enzymes again. The prefix and suffix are what allow multiple BioBricks to be combined easily. Outside of these parts, the BioBrick also consists of a plasmid backbone with a replication origin and antibiotic resistance marker, two parts necessary for cloning the plasmid (and the BioBrick with it).
Function
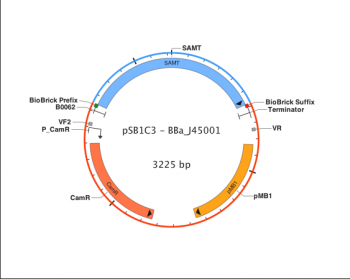
Each BioBrick has one function, and will complete only that function when combined with other BioBricks in a biological device. The function is often derived from the function of the DNA sequence the BioBrick was synthesized from. For instance, BBa_J45001, which encodes for a protein that converts salicylic acid to methyl salicylate (artificial wintergreen oil), is derived from Antirrhinus majus, a plant that naturally produces said protein. The most common way of finding proteins such as this is by looking for the desired function in various organisms and identifying the DNA sequence that causes the function. Then, the BioBrick is created by refining the DNA sequence in such a way that it only includes the desired function, and will not be affected by other biological parts or devices.
iGEM Registry
The iGEM registry collects the BioBricks and biological devices into one catalog from which iGEM teams and synthetic biology labs can order the parts they need. The catalog is organized online by type and function, with the following main categories:
- Promoters
- Ribosome Binding Sites
- Coding Sequences
- Terminators
There are also ways to organize by plasmid backbones or function. Each BioBrick has four pages related to it:
- Main Page: Gives a basic overview of how the BioBrick is used and how it functions
- Design: Information about the DNA sequence of the BioBrick and the plasmid, the ability of the BioBrick to be combined with different restriction enzymes, notes about the design process, and the source of the DNA sequence
- Experience: Allows users to submit their experiences using the BioBrick in their own biological devices
- Information: Technical information about the BioBrick itself, such as the original designer, part type, status, qualitative experience (whether it has worked in the past or not), and short description of its function
There is also a “part tools” section which includes related parts, the length of the plasmid, and sequence analyses, as well as ways to edit parts of the page. Finally, there is a link to order the part (assuming it is available).[5]
Future Directions
The iGEM Registry is a constantly updating resource, so it requires a lot of upkeep to run smoothly. There are many problems or missing information on the various pages of specific BioBricks, and the website is often undergoing changes to make finding and ordering the parts smoother. Most recently, the website was redesigned to simplify the page for each part, giving it just five individual pages to work with, while expanding the information available for it.[5] The iGEM Registry will continue to be refined and improved while expanding by the various additions from iGEM teams and synthetic biology labs around the world.
References
- Canton, B., Labno, A., and Endy, D. (2008) Refinement and standardization of synthetic biological parts and devices. Nature Biotechnology 26:787-793.
Discussion of the purpose and ideas behind designing biological parts and devices. - Knight, T., "Idempotent Vector Design for Standard Assembly of BioBricks" (MIT, Cambridge, MA, 2003).
Explanation of how and why BioBricks are designed and used. - Endy, D. (2005) Foundations for engineering biology. Nature 438:449-453.
Introduction and overview of the principles and ideas of synthetic biology. - Ho-Shing, O., Lau, K., Vernon, W., Eckdahl, T., Cambell, A. (2012) Assembly of standardized DNA parts using BioBrick ends in E. coli. Methods in Molecular Biology 852:61-76.
General methods and basic usage of BioBricks. - Registry of Standard Biological Parts. [1]
Online catalog of standard biological parts. Can be updated with new biological parts that fit the criteria.