CH391L/S14/Directed protein evolution
Contents |
What is directed protein evolution?
Evolution in nature consists of cycles of mutagenesis of an organism's DNA, environmental selection of the most fit mutants, and amplification of those favorable mutations by reproduction. This evolutionary principle has been "successfully exploited by humans over millennia to breed plants and animal." [1] However, more recently, researchers have utilized this evolutionary principle to evolve proteins at the molecular level, rather than evolving entire organisms. This method is referred to as directed protein evolution. Directed protein evolution is a powerful tool that is able to either optimize preexisting proteins or create novel proteins altogether. By developing a strong selective force, proteins are able to exhibit their plasticity by adapting to the environmental conditions set in a lab. By using directed protein evolution, a researcher does not need to completely understand the underpinnings of protein structures and various folds. Instead, iterative rounds of mutations and artificial selection generates proteins with desirable functions.
Process
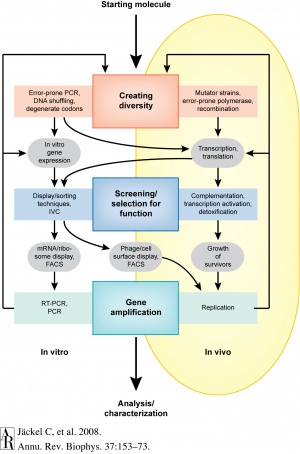
Each round of directed protein evolution consists of three steps: generating large libraries of randomly mutagenated copies of the gene of interest, appropriately selecting and screening for the desired phenotype or for a particular function, and amplifying the selected genes [1].
Library construction
Because we don't have nature's advantage of millions of years, we must artificially create large libraries of randomly mutagenated copies of the genes of interest. Common methods used for library construction include error-prone PCR and DNA shuffling.
Error-Prone PCR involves adding agents that increase the error-rate of DNA polymerase in PCR. Error-prone PCR methods generally include a higher concentration of MgCl2, which helps stabilize the non-complementary pairs [3]. Other common methods to increase the error-rate of DNA polymerase are adding Mn2+ and varying the ratio of nucleotides in the reaction. The previously explained methods have been able to increase the error-rate of DNA polymerase from 0.11% to 2% [3]. A disadvantage of error-prone PCR, however, comes from the fact that very few mutations are beneficial mutations. As a result, only single beneficial mutants are added during every generation of protein evolution.
DNA Shuffling circumvents the issue presented in error-prone PCR. First, a library of a gene of interest is created by error-prone PCR. Once the desired functions are screened for, the DNA of these clones are shuffled together to amass a large number of beneficial mutations. As these clones are bred iteratively, the frequency with which dramatic phenotype improvements occur increases dramatically, compared to error-prone PCR alone [3].
Screening and selection
After a sizable library of DNA is constructed, they must be expressed, either in vivo or in vitro, and evaluated for their ability to perform a particular function.This portion is the greatest challenge in directed protein evolution and lies at the crux of the experiment. The protein engineer is able to either select or screen for a desirable trait. The major difference of a selection and a screen is the fact that a selection simply gets rid of all of the variants that do not express a certain protein through the use of selective agents, such as antibiotics. Screens, however, are hand-picked variants that express a desired level of expression or another specific trait. To select for these desirable functions, high-throughput assays are used. To develop a high-throughput assay, there are two major hurdles that a protein engineer must navigate. First, the proteins that are being assayed for must be linked to the DNA that codes the polypeptide sequence. This is because DNA is much easier to isolate, sequence, and amplify. Second, a high-throughput assay must be developed that is compatible with the connection between protein and DNA. There are several main methods for tagging the proteins with DNA.
Physical Linkage Method simply creates a physical link between the protein and the DNA that encodes it. Several tactics used to create this physical link include phage display, ribosome display, peptide on plasmid, and cell surface display [4].
Compartmentalization Method restricts each protein and its associated DNA component into distinct compartments. This method of linking the genotype to the phenotype works especially well with assays that are based on enzyme catalysis. These assays include cell based assays and liposome-based assays [4].
Spatially Addressable Methods link the identity of a protein to a specific address in space. This way, if a desirable function of a protein is identified, the particular address in space can be linked back to the DNA sequence that encodes it. These assays include Microtiter-plate assays and protein chips [4].
Amplification
Once a variant is selected, in vitro methods, such as PCR, or in vivo methods, which simply allow the colonies to propagate, are used to amplify the desired gene.
Applications of Directed Protein Evolution
- Develop novel proteins that can perform functions outside of the context of the cell's survival
- Optimize the function of proteins
- study mechanisms of adaptions and protein structures [8]
Novel Methods for Directed Evolution
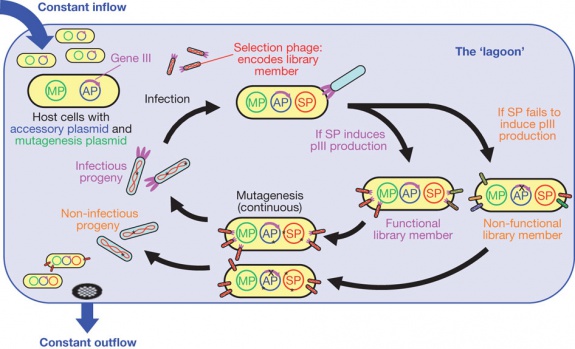
Although the methods mentioned have proved to be successful, a huge limitation is the amount of time that it takes for each step of the process. The method above is very time-consuming and requires frequent human intervention. As a response, a novel method of continuous protein evolution was developed, PACE (phage-assisted continuous evolution).
A lagoon of bacteriophage are present and E.coli are moved through this "lagoon" at a rate faster than which they can divide, but long enough for the phage to infect them and divide. These phages contain the gene of interest. To preface, phages require the pIII gene in order for it to take over the host cell, replicate, and eventually lyse. Experimentalists deleted this gene from the phages' genome and placed it into an accessory plasmid located in the host cells. Upstream from the pIII gene in the accessory plasmid, there is a selective agent (i.e promoter sequences, protein-protein recognition, etc). Therefore, if the gene product of the phage is able to induce the expression of the pIII gene in the accessory plasmid, the phages replicate, lyse the cell, and reenter the "lagoon". If the phages' gene product does not induce the upstream promoter, however, the phage does not replicate and is instead flushed away with the other bacteria that didn't successfully express the pIII gene. The mutagenesis comes from an arabinose-inducible mutagenesis plasmid; it elevates the error rate by suppressing proofreading and enhancing error-prone lesion bypass [2].
Future Direction
- Further investigation of the evolution of proteins will give insight to protein engineers who seek to rationally design proteins
- Quicker and more accurate screenings
- Engineering networks of interacting proteins
References
- Christian Jäckel, Peter Kast, and Donald Hilvert. Protein Design by Directed Evolution. Annual Review of Biophysics, 2008.
- Kevin M. Esvelt, Jacob C. Carlson & David R. Liu. A system for the continuous directed evolution of biomolecules. Nature 472, 499–503 (28 April 2011) doi:10.1038/nature09929
- Ling Yuan1, Itzhak Kurek, James English and Robert Keenan. Laboratory-Directed Protein Evolution. Microbiol. Mol. Biol. Rev. September 2005 vol. 69 no. 3 373-392.
- Lin, H. and Cornish, V. W. (2002), Screening and Selection Methods for Large-Scale Analysis of Protein Function. Angew. Chem. Int. Ed., 41: 4402–4425. doi: 10.1002/1521-3773(20021202)41:23<4402::AID-ANIE4402>3.0.CO;2-H
- Giver L, Gershenson A, Freskgard PO, Arnold FH. Directed evolution of a thermostable esterase. Proc Natl Acad Sci USA, 1998.
- Patnaik, R., S. Louie, V. Gavrilovic, K. Perry, W. P. Stemmer, C. M. Ryan, and S. del Cardayre. 2002. Genome shuffling of Lactobacillus for improved acid tolerance.
- Song, J. K., B. Chung, Y. H. Oh, and J. S. Rhee. 2002. Construction of DNA-shuffled and incrementally truncated libraries by a mutagenic and unidirectional reassembly method: changing from a substrate specificity of phospholipase to that of lipase. Appl. Environ. Microbiol. 68:6146-6151
- Kuhlman, B., G. Dantas, G. C. Ireton, G. Varani, B. L. Stoddard, and D. Baker. 2003. Design of a novel globular protein fold with atomic-level accuracy. Science 302:1364-1368
- Matthew W. Peters, Peter Meinhold, Anton Glieder,and Frances H. Arnold†, Regio- and Enantioselective Alkane Hydroxylation with Engineered Cytochromes P450 BM-3.
- Romero PA and Arnold FH. Exploring protein fitness landscapes by directed evolution. Nat Rev Mol Cell Bio, 2009.