CH391L/S14/SelectableCounterselectable
Contents |
Selectable and Counter-Selectable Genetic Markers
Selectable and Counter-Selectable Markers are a fundamental part of synthetic biology. Isolation of cells containing certain alleles is a vital part of many experiments. For instance, currently there exists no method of genetic transformation in which 100% of cells that are exposed to the exogenous DNA will take up and incorporate it, so that it may be transcribed into RNA (and then translated into proteins). Thus, selection is essential to synthetic biology, as it allows for the percentage of cells that have incorporated DNA into their genetic code to be selected. Counter selection too has many important applications, such as isolating mutants, or performing dual selection (in conjunction with selection processes).
Selectable Genetic Markers
Selectable Genetic Markers are markers that are added to a genetic sequence that allows for the host cell to survive in a certain environment that would be lethal to cells that do not have the markers. Thus, selectable genetic markers are used to induce positive selection (aka “selection”) a process in which cells that possess certain, favorable traits that allow them to survive in a certain environment are isolated from those that do not possess the trait. One could, for instance, enable a cell to be resistant to an antibiotic via the insertion of a marker that enables the cell to develop immunity to that antibiotic. The cell culture can then be exposed to the antibiotic, and cells that possess the marker would be able to survive, while the cells that do not are unable to survive. In this case, the culture underwent positive selection in regards to the antibiotic-resistant marker gene.
Example of a Selectable Marker
Hygromycin Phosphotransferase (hph): Derived from E.Coli, the marker encodes for a resistance to aminocyclitol antibiotic hygromycin B. Hygromycin B inhibits protein synthesis by “interfering with ribosomal translocation and with aminoacyl-tRNA recognition.” [1] This marker is especially useful due to the fact that there are no cell lines that are naturally resistant to Hygromycin B. This allows the antibiotic to be used in selection (specifically positive selection) when the hph marker gene is inserted into the cell. Furthermore, a study done in 1984 showed that this marker gene is useful as it can be used in eukaryotic cell lines (however pMLTR-I needs to be used as a vector in addition to the hph gene for eukaryotic cell lines). In general, the marker is very effective, experiments involving positive selection with the marker produce very few false positives, and is today the most widely used selectable markers in genetic manipulation of Histoplasma capsulatum. [1] [2]
Counter Selectable Genetic Markers
Counter Selectable Genetic Markers are markers that are added to the genetic sequence that are either toxic to the cell or inhibitory to cell replication. These elements are often incorporated into genetic modification schemes in order to select for rare recombination events that require the removal of the marker or to selectively eliminate plasmids or cells from a given population. Counter Selectable markers are thus used to induce negative selection, a process in which unwanted elements are eliminated from the population. Negative selection can occur without the use of counter selectable markers, as it could occur by exploiting traits that are naturally found in certain cell lines. In a population of T cells, B cells, monocytes, and NK cells, introducing antibodies specific to B cells, NK cells, and monocytes is one method of performing negative selection against those cell types to isolate T cells. [3]
Counter selectable markers are used in either dual selection or in isolating cells that mutated in such a way to to render the marker’s inhibitory or lethal allele ineffective or not present.
Examples of Counter-Selectable Markers
Tdk genetic marker: This gene acts as a counter selectable marker in the presence of AZT (a chain terminating base analogue). When incubated in media containing AZT, cells that incorporated the tdk gene into their genetic code are killed by AZT. For dual selection purposes, the tdk gene is often used in conjunction with the Kan gene, which confers survival against Kanamycin. As with many counter selectable markers, this marker produces a “very noisy and subtle selection,” with no known reason as to why (Metzgar et al speculate that this is either due to “the low efficiency of the tdk construct used” or “a reduction in the toxicity of chain termination in the presence of the Acinetobacter ADP1 recombination machinery”). In general, though, the majority of the population will be tdk– mutants after negative selection in the presence of AZT. [4]
False negatives are a problem in most genes used for counter selection. This issue is addressed (and, for the most part, resolved) by Barkan et al by using galK and 2-deoxy-galactose in addition to sacB.
galK and 2-deoxy-galactose (used in conjunction with sacB) – A widely used counter-selectable gene encodes for sacB, which makes sucrose toxic to the cell and thus can be used as a counter selectable marker. This marker, however, frequently undergoes spontaneous inactivation, which hinders its ability to act as a counter selectable marker. This problem can be solved by the introduction of the galK marker and 2-deoxy-galactose in addition to the sacB marker vastly reduced the amount of mutational inactivation in both markers (to a point where the mutational inactivation was lower then the recombination requency). [5]
TolC - An outer membrane pore thats found in E.coli that is involved in transmembrane transport and has been implemented in selection and counter selection. Usually the escape frequency for counter selection with colicin (a type of toxin produced by E. Coli) is too high for TolC to be used as a counter-selectable marker. By introducing vancomycin, the escape frequency for counter selection was reduced by 425 fold. The study also was able to create a strain of E. coli that is optimal for genome engineering as it is roughly 10 times more efficient in achieving allelic diversity [6].
Dual Selection and Screening
Dual Selection - A process in which both positive and negative selection are implemented (often cycles of positive selection followed negative selection). Often this is done by the use of both selectable and counter selectable markers (in which the selectable marker's allele is exploited for positive selection, and the counter selectable marker's allele is exploited for negative selection).
Screenable Genetic Markers are vectors that are added to the genetic sequence that code for a protein that is neither detrimental nor helpful to the cell, but rather expresses a distinct property that allows for the observation of how many cells have integrated the synthetic DNA sequence (this is often done by coding for a fluorescent protein such as GFP).
Screening Equipment
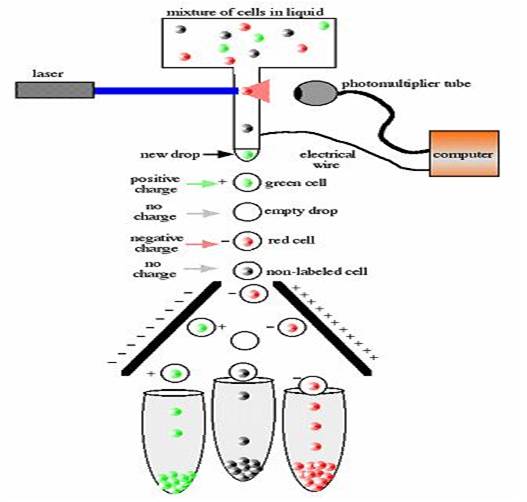
Flow Cytometer - Used for screening purposes, this machine sorts individual cells by detecting screenable genetic markers. It makes it possible to sort bacteria that are diluted in liquid, as opposed to being plated on solid gel. It does this by passing droplets, containing only one bacterium per droplet, through a laser that is pointed at a sensor that detects fluorescence. The sensor senses what light wave the cell emits and electrically charges the droplet (either positive or negative), or in the case of the cell not emitting light, the cytometer places no charge on the droplet. The droplet then free falls through past two parallel, oppositely charged plates. Depending on the charge that was placed on the droplet, it would either be attracted to the positively charged plate, negatively charged plate, or neither. Three collection tubes are placed at the bottom of the cytometer. The droplet that were attracted to the positively charged plate go in one tube, the ones that weren't attracted to either plate go in the center tube, and the droplets that were attracted to the negatively charged plate fall in the other tube.
Examples of Dual Selection
Selection and Screening of Gene Switches in Escherichia coli
The study describes a technique to identify bacteria that exhibit a genetic switch, which they liken to a switch in an electric circuit. The presence of this switch would either turn on or turn off an expression of an allel, such as antibiotic resistance or salt sensitivity. They used a library of genetic traits (or 'riboswitches') to investigate, and the technique that they used involved attaching a fluorescent dye to the 'switch.'
They then used a flow cytometer to sort out those bacteria having the 'switch' from those not having it. The bacteria that have the 'switch' will fluoresce while those without will not.
Genetic switches are proteins that bind to a sequence of DNA and alter its expression. Depending on certain, specified conditions (usually the presence, or lack, of a chemical or physical signal), these switches can either be set to “ON” or “OFF.” Duel Selection can be implemented for ensuring the expression of genetic switches. Positive selection markers that are expressed in conditions that induce the gene switch to be ON and negative selection markers that are expressed when such conditions cause the gene switch to be OFF are both implemented. Through cycles of positive and negative selection, the cells that code for the switch are isolated. This method, however, has problems as it requires plasmid isolation steps and increases the likelihood of false positives.
tetA - “encodes a tetracycline/H+ antiporter as both a positive and a negative selection marker.” The protein allows for dual selection without the need to introduce multiple vectors. “Expression of TetA confers the cells tetracycline resistance (ON selection) while the overexpression of the membrane-bound protein renders them more sensitive to toxic metal salts such as NiCl2 as well as other compounds (OFF selection).” [7] This process produces fewer false positives, and makes selection easier.
Liquid bacterial cultures present a problem for selection markers, as false positive clones are difficult to isolate and in some cases the false positive could have a shorter replication time, and could take over the colony. Muranaka et al says the group anticipates the tetA selection marker could effectively be used in liquid cultures as it produces far fewer false positives. The group tested the efficiency of the dual selection marker by running a series of mock selections using riboswitches and model background clones. Their library consisted of four TPP riboswitch phenotypes: an OFF switch (–ThiMwt), an ON switch (+ThiM#2), an ALWAYS ON switch (ThiM-ON), and an ALWAYS OFF switch (ThiM-OFF). The group ran this trial in liquid media and used flow cytometry to count and sort the bacteria cells that were fluorescent. Though the method could be used for both solid and liquid media, after running the colony through positive and negative selection the group determined that the selection was optimized when the bacteria was in liquid culture at high concentrations. The selection is also useful as the host cell does not need to have a specific phenotype unless in the circumstance of there being an extrachromosomal element that allow for tetracycline resistance. [7]
Enhancing Signaling Specificity of Transcriptional Activator LuxR
Transcriptional factor LuxR activates when bound to the signaling molecule 3OC6HSL, “an acyl-HSL with a carbonyl substituent at the third carbon of the acyl chain.” Other variants of this transcriptional factor exist, such as the LuxR-G2E variant, that respond to different acyl chains. Collins et al implements directional evolution to produce a variety of mutated LuxR transcriptional factors, then implements a dual selection system to identify a variant of LuxR that still responds to straight-chain acyl-HSLs, but has no response to 3OC6HSL. [8]
LuxR, and mutants of LuxR, are genetic switches. Dual selection, as described in the previous section, is a useful method of selection available for genetic switches. In this case, the ON selection plasmid is pluxCAT (which enables the bacteria to be resistant to chloramphenicol), while the OFF selection plasmid codes for pluxBLIP (which encodes Beta-lactamase inhibitory protein). After each round of selection, both pluxCAT and pluxBLIP are deactivated by cleavage with a restriction endonuclease. The reporter plasmid pluxGFPuv, which contains the screening marker GFPuv which can be activated by the lux promoter, is used to assess the how the mutants of the LuxR genetic switch respond to various acyl chain molecules. Introducing straight-chain acyl-HSLs to the bacteria in the presence of chloramphenicol induced positive selection. For the negative selection run, 3OC6HSL, C10HSL, C6HSL and C8HSL were introduced into the media along with carbenicilin. The cells possessing the variant LuxR-G2E had the genetic switch OFF and thus resisted the carbenicilin (due to the OFF selection plasmid coding for pluxBLIP). Thus, LuxR-G2E a more signal-specific version of LuxR, was isolated. [8]
iGEM and Future Directions
In 2012, the Edinburgh team used sucrose, instead of an antibiotic, as an effective tool for selection. Sucrose hydrolase was used as the selective marker, while nitroreductase and SacB were used for counter selection markers. Using sucrose instead of an antibiotic as a means of providing an environment with which cells could undergo selection and counter selection is helpful, as bacteria that are resistant to antibiotics have the potential to cause an epidemic of widespread disease (which it is against international law to release genetically modified bacteria with such resistance to bacteria into the wild). [9]
Articles regarding selection and counter-selection are currently mainly focused on reducing the number of false positives, and reducing the amount of vectors needed to reach a desired result. In the future, it is likely this trend would continue, however the Edinburgh team does make a point as to the safety regarding these bacteria, and further research on using the use of sucrose as a means of selection ought to be done.
An article published by Herbert Schweizer in 2008 seems to suggest that the future of selection is in methods that do not require antibiotic resistance, but rather by methods that use chemicals such as sucrose as a mechanism for selection. The paper also talks about the growing use of counter-selection as a means of isolating a property and implies that this method will be implemented more in the future. [10]
References
- K. Blochlinger "Hygromycin B phosphotransferase as a selectable marker for DNA transfer experiments with higher eucaryotic cells" (1984 American Society for Microbiology).
Information regarding hph and uses in eukaryotic cell lines - A. Smulian "Expression of Hygromycin Phosphotransferase Alters Virulence of Histoplasma capsulatum" (2007 American Society for Microbiology).
Information regarding hph and Histoplasma capsulatum - “http://technical.sanguinebio.com/positive-selection-vs-negative-selection-for-cell-isolation/”
Example of isolating T cells using counter selection - D. Metzgar, “Acinetobacter sp. ADP1: an ideal model organism for genetic analysis and genome engineering” (2004 Nucleic Acids Res)
Information regarding the tdk selective marker - D. Barkan, "An improved counterselectable marker system for mycobacterial recombination using galK and 2-deoxy-galactose" (2010 Elsevier B.V.).
Information regarding galK and 2-deoxy-galactose used with sacB - J. Gregg, "Rational optimization for tolC as a powerful dual selectable marker for genome engineering"
Information regarding tolC and the use of vancomycin to improve its selection capabilities - N. Muranaka, "An efficient platform for genetic selection and screening of gene switches in Escherichia coli" (2009 Nucleic Acid Res.).
Information regarding tetA and its use in dual selection of gene switches (specifically the TPP Riboswitch) - CH. Collins "Dual selection enhances the signaling specificity of a variant of the quorum-sensing transcriptional activator LuxR."
Information regarding the study that isolated LuxR-G2E using dual selection - iGEM "http://2012.igem.org/wiki/index.php?title=Team:Edinburgh/Project/Non-antibiotic-Markers"
Information regarding the Edinburgh team’s implementation of sucrose as an agent for selection - H. Schweizer "Bacterial genetics: past achievements, present state of the field, and future challenges" (2008 Biotechniques).
Information regarding the future of selectable and counter selectable markers